Unconstrained optimization - EGO#
[1]:
import numpy as np
from matplotlib import pyplot as plt
from smt_optim.benchmarks.registry import get_problem
from smt_optim.core import Driver, ObjectiveConfig, ConstraintConfig, DriverConfig, Problem
from smt_optim.surrogate_models.smt import SmtAutoModel
from smt_optim.acquisition_strategies import MFSEGO
Unconstrained 1D optimization#
This example will use the Sasena test function (2002). The bounds are \([0, 10]\)
[2]:
def sasena_2002(x: np.ndarray):
return -np.sin(x) - np.exp(x / 100) + 10
bounds = np.array([[0, 10]])
[3]:
obj_config = ObjectiveConfig(
objective=[sasena_2002],
type="minimize",
surrogate=SmtAutoModel, # set which GP to model the objective
)
prob_definition = Problem(
obj_configs=[obj_config],
design_space=bounds,
)
Once the problem is initialized, we can configure the optimization driver. We can set the configuration using the DriverConfig dataclass and then initialize the driver with the Driver class.
In our example, we set the following parameters:
max_itercorresponds to the maximum number of iteration that will be performed.nt_initcorresponds to the number of samples in the initial DOE. The initial DOE will be generated using LHS.verbosecan be set toTrueorFalse. If set toTruesome basic information will be printed at the end of each iteration.seedallows to reproduce optimization run.
The MFSEGO optimization strategy must be pass to perform EGO (unconstrained optimization), SEGO (constrained optimization) and MFSEGO (unconstrained or constrained and multi-fidelity optimization).
[4]:
driver_config = DriverConfig(
max_iter = 5, # stopping criterion
nt_init = 2, # number of sample in the initial DoE
verbose = True,
seed=42,
)
# configure the optimizer. Note: if a single fidelity level is given to the MFSEGO acquisition strategy, it falls back to the SEGO framework.
driver = Driver(prob_definition, driver_config, MFSEGO)
We can launch the optimization using the optimize() class method. It will return a State object.
[5]:
# return the optimization data
state = driver.optimize()
iter budget fmin fidelity gp_time acq_time
1 3 8.13516e+00 1 0.046 0.065
2 4 8.11303e+00 1 0.127 0.067
3 5 8.03882e+00 1 0.138 0.047
4 6 7.91831e+00 1 0.093 0.047
5 7 7.91826e+00 1 0.091 0.046
Once the optimization terminated, we can recover the best sample using the get_best_sample() class method. It will return a Sample dataclass with the lowest objective value.
[6]:
sample = state.get_best_sample()
sample
[6]:
======= sample data =======
x = [7.85760188]
obj = [7.91826097]
cstr = []
eval_time = [6.29399437e-06]
------- meta data -------
iter = 5
budget = 7
fidelity = 0
===========================
We can only also recover the entire DOE using the dataset attribute.
[7]:
dataset = state.dataset
data = dataset.export_as_dict()
xt = data["x"]
yt = data["obj"]
[8]:
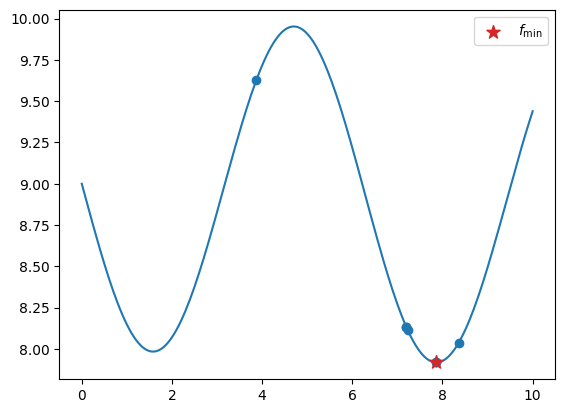
x_test = np.linspace(0, 10, 201)
y_test = sasena_2002(x_test)
fig, ax = plt.subplots()
ax.plot(x_test, y_test)
ax.scatter(xt, yt)
ax.scatter(sample.x, sample.obj, 100, marker="*", color="C3", zorder=20, label=r"$f_\min$")
ax.legend()
plt.show()
Unconstrained 2D optimization#
[9]:
def modified_branin(x):
"""
Engineering Design via Surrogate Modelling: A Practical Guide
A. I. J. Forrester, A. Sóbester and A. J. Keane
2008
"""
X1 = 15 * x[0] - 5
X2 = 15 * x[1]
a = 1
b = 5.1 / (4 * np.pi ** 2)
c = 5 / np.pi
d = 6
e = 10
ff = 1 / (8 * np.pi)
f = (a * (X2 - b * X1 ** 2 + c * X1 - d) ** 2 + e * (1 - ff) * np.cos(X1) + e) + 5 * x[0]
return f
bounds = np.array([
[0, 1],
[0, 1],
])
[10]:
obj_config = ObjectiveConfig(
[modified_branin],
type="minimize",
surrogate=SmtAutoModel,
)
prob_definition = Problem(
obj_configs=[obj_config],
design_space=bounds, # problem bounds
)
opt_config = DriverConfig(
max_iter = 20,
nt_init = 3,
verbose = True,
scaling = True,
seed=42,
)
driver = Driver(prob_definition, opt_config, MFSEGO)
state = driver.optimize()
iter budget fmin fidelity gp_time acq_time
1 4 1.56584e+01 1 0.141 0.252
2 5 1.55312e+01 1 0.289 0.488
3 6 1.51473e+01 1 0.250 0.375
4 7 1.42022e+01 1 0.271 0.340
5 8 6.94080e+00 1 0.273 0.199
6 9 6.94080e+00 1 0.236 0.251
7 10 3.13434e+00 1 0.283 0.295
8 11 1.85202e+00 1 0.257 0.381
9 12 1.85202e+00 1 0.308 0.392
10 13 1.85202e+00 1 0.316 0.496
iter budget fmin fidelity gp_time acq_time
11 14 1.01441e+00 1 0.313 0.470
12 15 1.01260e+00 1 0.345 0.464
13 16 1.01260e+00 1 0.277 0.377
14 17 1.01157e+00 1 0.299 0.546
15 18 1.01157e+00 1 0.269 0.851
16 19 1.01157e+00 1 0.244 0.948
17 20 1.01157e+00 1 0.234 0.733
18 21 1.01157e+00 1 0.211 0.856
19 22 1.01157e+00 1 0.263 0.715
20 23 1.01157e+00 1 0.250 0.822
[11]:
# get the best sample in the dataset
sample = state.get_best_sample()
xt = state.dataset.export_as_dict()["x"]
X = np.linspace(bounds[0, 0], bounds[0, 1], 201)
Y = np.linspace(bounds[1, 0], bounds[1, 1], 201)
XX, YY = np.meshgrid(X, Y)
data = np.vstack((XX.ravel(), YY.ravel())).T
z = np.empty(data.shape[0])
for i in range(data.shape[0]):
z[i] = modified_branin(data[i, :])
Z = z.reshape(XX.shape)
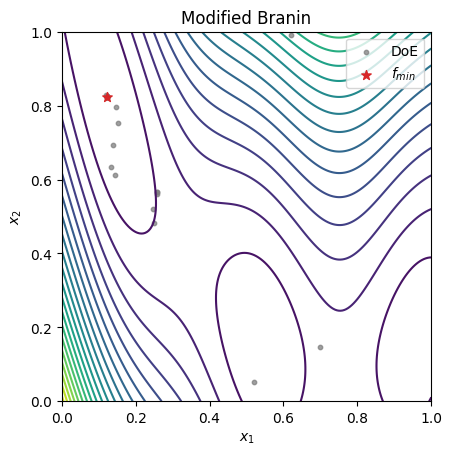
fig, ax = plt.subplots()
ax.set_title("Modified Branin")
# plot the test function
ax.contour(XX, YY, Z, levels=20)
# plot the points in the final DoE
ax.scatter(xt[:, 0], xt[:, 1], 10, color="C7", alpha=0.75, label="DoE")
# plot the best sample in the final DoE
ax.scatter(sample.x[0], sample.x[1], 50, c="C3", marker="*", label=r"$f_{min}$", zorder=30)
ax.set_xlabel(r"$x_1$")
ax.set_ylabel(r"$x_2$")
ax.legend()
ax.set_aspect("equal")
plt.show()