Mixed-variable optimization#
This page demonstrates how to minimize mixed-variable problems. SMT Optim supports float, integer and categorical variables.
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import proj3d
# mixed-variables optimization requires the use of SMT's DesignSpace class
import smt.design_space as ds
from smt_optim import minimize
# method to fetch optimization problems
from smt_optim.benchmarks.registry import get_problem
# utility method to help with plotting 2d functions
from smt_optim.utils.plot_2d import get_plot2d_data
Unconstrained mixed-variable optimization#
Defining the test problem#
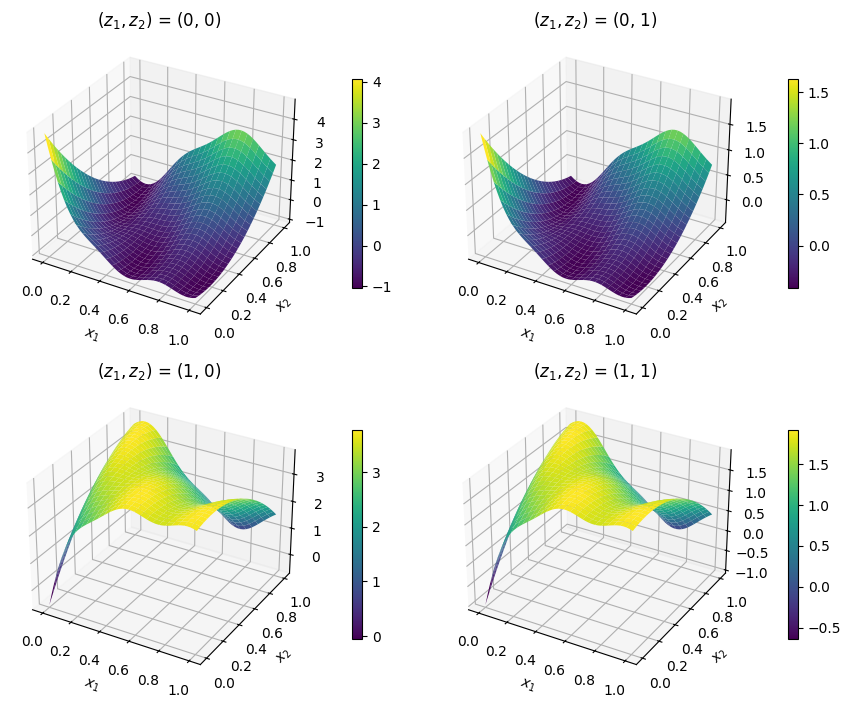
The following example illustrates how to minimize the mixed-variable Branin test function. This example assumes the user is familiar with unconstrained optimization of a continuous design space. This problem has 2 continuous ‘float’ variables (\(x_1,\; x_2\)) and 2 categorical variables (\(z_1,\; z_2\)). It can be written mathematically as:
The goal is to minimize the objective function over the mixed-variable design space, which can be formulated as so:
The cell below imports the test function from within SMT Optim test problems collection. The mixed-variable design space is initialized with an SMT’s DesignSpace class.
# fetches the mixed-variable test problem
problem = get_problem("MixVarBranin")
# defines the design space using SMT's DesignSpace class
design_space = ds.DesignSpace([
ds.FloatVariable(lower=0, upper=1),
ds.FloatVariable(lower=0, upper=1),
ds.CategoricalVariable([0, 1]),
ds.CategoricalVariable([0, 1]),
])
# ------- plots the test problem -------
bounds = np.array([
[0, 1],
[0, 1],
])
z_combinations = [
(0, 0),
(0, 1),
(1, 0),
(1, 1),
]
fig = plt.figure(figsize=(9, 7))
for i, (z1, z2) in enumerate(z_combinations, start=1):
def wrapper(x):
x0 = np.array([x[0], x[1], z1, z2])
return problem.objective(x0)
XX, YY, Z = get_plot2d_data(wrapper, bounds, 51)
ax = fig.add_subplot(2, 2, i, projection="3d")
surf = ax.plot_surface(XX, YY, Z, cmap="viridis", edgecolor="none")
ax.set_title(r"$(z_1, z_2)$ = (" + str(z1) + ", " + str(z2) + ")")
ax.set_xlabel(r"$x_1$")
ax.set_ylabel(r"$x_2$")
fig.colorbar(surf, ax=ax, shrink=0.7, pad=0.1)
plt.tight_layout(h_pad=2.5)
plt.show()
Starting the optimization#
The Bayesian optimization is started with the minimize method. To define a mixed-variable design space instead of a continuous one, an SMT’s DesignSpace object must be given to the design space argument.
# starts the optimization
state = minimize(
[problem.objective], # list of callables (single callable for mono-fidelity)
design_space, # SMT's Design Space object
max_iter=10,
driver_kwargs={"seed": 0}
)
# fetches the best sample
best_sample = state.get_best_sample()
print(best_sample)
iter budget fmin rscv fidelity gp_time acq_time
1 6 -6.95818e-01 0.000e+00 1 0.368 1.613
2 7 -6.95818e-01 0.000e+00 1 0.393 1.855
3 8 -6.95818e-01 0.000e+00 1 0.414 1.127
4 9 -9.02607e-01 0.000e+00 1 0.390 1.443
5 10 -9.02607e-01 0.000e+00 1 0.428 1.357
6 11 -9.52011e-01 0.000e+00 1 0.456 1.474
7 12 -1.04257e+00 0.000e+00 1 0.485 1.533
8 13 -1.04257e+00 0.000e+00 1 0.557 1.221
9 14 -1.04257e+00 0.000e+00 1 0.393 1.233
10 15 -1.04373e+00 0.000e+00 1 0.383 1.286
======= sample data =======
x = [0.55584733 0.14561377 0. 0. ]
obj = [-1.04372608]
cstr = []
eval_time = [1.02920021e-05]
------- meta data -------
iter = 10
budget = 15
fidelity = 0
rscv = 0.0
===========================
Plotting the results#
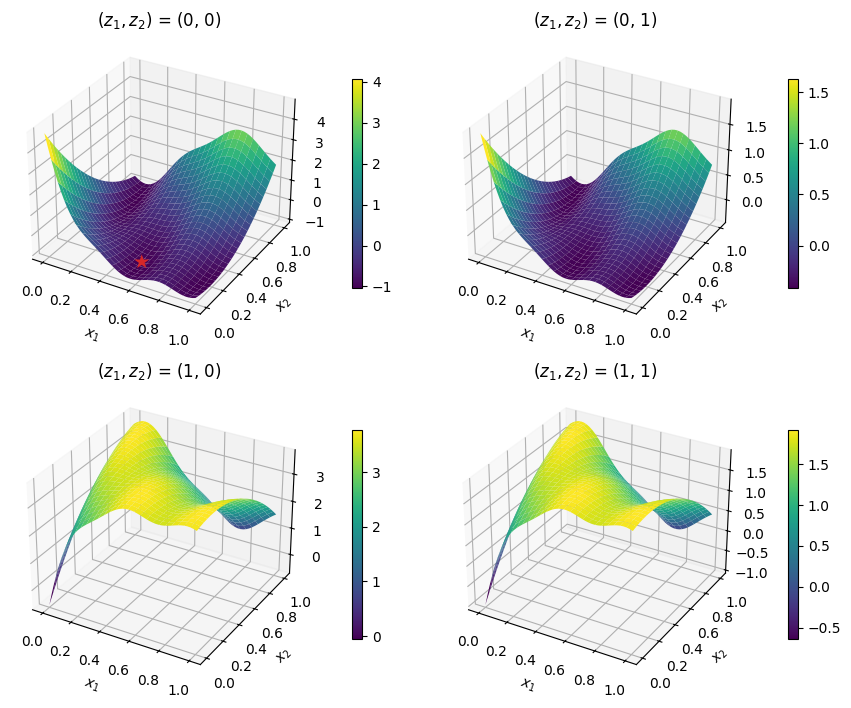
The code snippet below plots the best sample, displayed with a star, against the test problem.
x_star = best_sample.x
y_star = best_sample.obj[0]
# categorical variables
z1_star = x_star[2]
z2_star = x_star[3]
# ------- plots the best sample against the functions -------
fig = plt.figure(figsize=(9, 7))
for i, (z1, z2) in enumerate(z_combinations, start=1):
def wrapper(x):
x0 = np.array([x[0], x[1], z1, z2])
return problem.objective(x0)
XX, YY, Z = get_plot2d_data(wrapper, bounds, 51)
ax = fig.add_subplot(2, 2, i, projection="3d")
surf = ax.plot_surface(XX, YY, Z, cmap="viridis", edgecolor="none", zsort="min")
ax.set_title(r"$(z_1, z_2)$ = (" + str(z1) + ", " + str(z2) + ")")
ax.set_xlabel(r"$x_1$")
ax.set_ylabel(r"$x_2$")
fig.colorbar(surf, ax=ax, shrink=0.7, pad=0.1)
if z1 == z1_star and z2 == z2_star:
x2, y2, _ = proj3d.proj_transform(
x_star[0], x_star[1], y_star,
ax.get_proj()
)
ax.annotate(
"★",
(x2, y2),
color="C3",
fontsize=14,
ha="center"
)
plt.tight_layout(h_pad=2.5)
plt.show()
Constrained, multi-fidelity, and mixed-variable optimization#
Defining the test problem#
The following example illustrates how to minimize the constrained, multi-fidelity, mixed-variable Branin test function. This example assumes the user is familiar with constrained and multi-fidelity optimization. The problem’s design space has 2 continuous ‘float’ variables and 2 categorical variables. The mixed-variable constraint function is defined as:
The multi-fidelity objective and constraint functions take the following form:
where \(f(\boldsymbol x)\) is the same objective function as the previous example. The goal is to minimize the high-fidelity objective function with respect to the inequality constraint and leverage their ‘cheaper’ approximation to reduce the optimization cost.
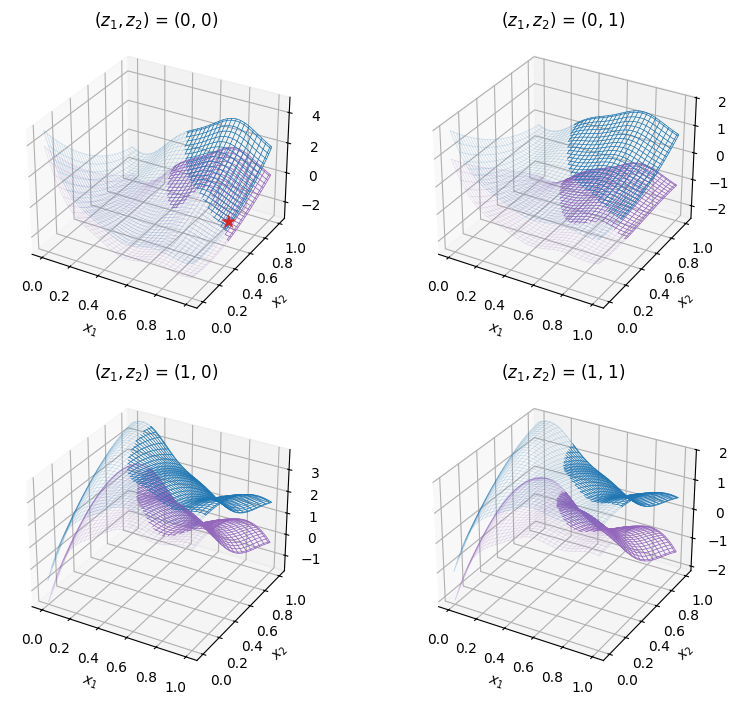
The cell below imports the test function from within SMT Optim test problems collection. The design space is initialized with an SMT’s DesignSpace class. The figure shows the low- (in purple) and high-fidelity (in blue) test problems over their entire design space. The semi-transparent regions denote the unfeasible regions due to the inequality constraint.
# fetches the mixed-variable test problem
problem = get_problem("MultiFidelityMixVarBranin")
# defines the design space using SMT's DesignSpace class
design_space = ds.DesignSpace([
ds.FloatVariable(lower=0, upper=1),
ds.FloatVariable(lower=0, upper=1),
ds.CategoricalVariable([0, 1]),
ds.CategoricalVariable([0, 1]),
])
# ------- plots the test problem -------
bounds = np.array([
[0, 1],
[0, 1],
])
z_combinations = [
(0, 0),
(0, 1),
(1, 0),
(1, 1),
]
colors = ["C4", "C0"]
fig = plt.figure(figsize=(9, 7))
for i, (z1, z2) in enumerate(z_combinations, start=1):
ax = fig.add_subplot(2, 2, i, projection="3d")
for lvl in range(problem.num_fidelity):
def wrapper(x, func):
x0 = np.array([x[0], x[1], z1, z2])
return func(x0)
XX, YY, Z = get_plot2d_data(lambda x, f=problem.objectives[lvl]: wrapper(x, func=f), bounds, 101)
_, _, G = get_plot2d_data(lambda x, f=problem.constraints[0][lvl]: wrapper(x, func=f), bounds, 101)
Z_feas = np.where(G > 0, np.nan, Z)
ax.set_title(r"$(z_1, z_2)$ = (" + str(z1) + ", " + str(z2) + ")", pad=5.0)
ax.set_xlabel(r"$x_1$")
ax.set_ylabel(r"$x_2$")
ax.plot_wireframe(XX, YY, Z_feas, linewidth=0.5, edgecolor=colors[lvl])
ax.plot_wireframe(XX, YY, Z, alpha=0.2, linewidth=0.5, edgecolor=colors[lvl])
plt.tight_layout(h_pad=2.5)
plt.show()
Starting the optimization#
The Bayesian optimization is started by calling the minimize method. To define a mixed-variable design space instead of a continuous one, an SMT’s DesignSpace object must be given to the design space argument. Additionally, the objective argument expects a list of method callables; the same applies to any constraints.
# defines the constraint
constraint = [
{
"fun": problem.constraints[0], # list of callables
"upper": 0., # g(x) <= 0
}
]
# starts the optimization
state = minimize(
problem.objectives, # list of callables
design_space, # ds.DesignSpace object
constraints=constraint,
costs=[0.2, 1.],
max_iter=10,
max_budget=10,
# arguments passed to the optimization driver
driver_kwargs={
"nt_init": 3,
"seed": 42
},
# arguments passed to the acquisition strategy
strategy_kwargs={
"fidelity_crit": "pessimistic",
"sp_method": "Cobyla",
"sp_tol": 1e-4
}
)
# fetches the best sample
best_sample = state.get_best_sample()
print(best_sample)
iter budget fmin rscv fidelity gp_time acq_time
1 4.400 2.70574e+00 0.000e+00 1 2.841 2.537
2 4.600 2.70574e+00 0.000e+00 1 1.602 2.157
3 4.800 2.70574e+00 0.000e+00 1 1.660 1.996
4 5.000 2.70574e+00 0.000e+00 1 1.579 2.423
5 5.200 2.70574e+00 0.000e+00 1 1.252 2.725
6 5.400 2.70574e+00 0.000e+00 1 1.644 4.044
7 5.600 2.70574e+00 0.000e+00 1 1.657 15.491
8 6.800 -7.36781e-01 0.000e+00 2 1.374 5.550
9 7.000 -7.36781e-01 0.000e+00 1 1.458 3.358
10 7.200 -7.36781e-01 0.000e+00 1 1.751 3.047
======= sample data =======
x = [1. 0.43797134 0. 0. ]
obj = [-0.73678137]
cstr = [-0.03797134]
eval_time = [2.54299084e-06 6.27012923e-07]
------- meta data -------
iter = 8
budget = 6.800000000000002
fidelity = 1
rscv = 0.0
===========================
Plotting the results#
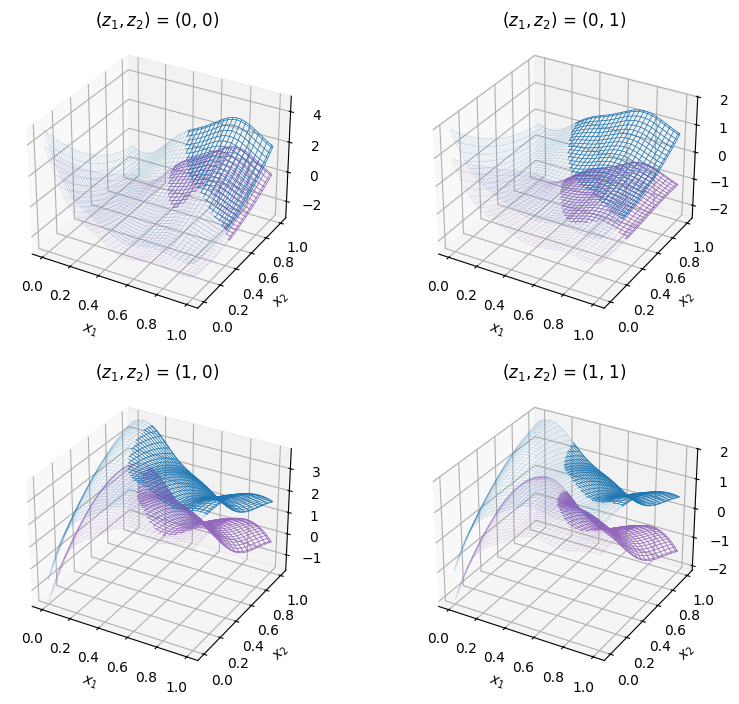
The code snippet below plots the best sample, displayed with a star, against the test problem.
best_sample = state.get_best_sample()
x_star = best_sample.x
y_star = best_sample.obj[0]
# categorical variables
z1_star = x_star[2]
z2_star = x_star[3]
# ------- plots the best sample against the functions -------
fig = plt.figure(figsize=(9, 7))
for i, (z1, z2) in enumerate(z_combinations, start=1):
ax = fig.add_subplot(2, 2, i, projection="3d")
for lvl in range(problem.num_fidelity):
def wrapper(x, func):
x0 = np.array([x[0], x[1], z1, z2])
return func(x0)
XX, YY, Z = get_plot2d_data(lambda x, f=problem.objectives[lvl]: wrapper(x, func=f), bounds, 101)
_, _, G = get_plot2d_data(lambda x, f=problem.constraints[0][lvl]: wrapper(x, func=f), bounds, 101)
Z_feas = np.where(G > 0, np.nan, Z)
ax.set_title(r"$(z_1, z_2)$ = (" + str(z1) + ", " + str(z2) + ")")
ax.set_xlabel(r"$x_1$")
ax.set_ylabel(r"$x_2$")
ax.plot_wireframe(XX, YY, Z_feas, linewidth=0.5, edgecolor=colors[lvl])
ax.plot_wireframe(XX, YY, Z, alpha=0.2, linewidth=0.5, edgecolor=colors[lvl])
if z1 == z1_star and z2 == z2_star:
x2, y2, _ = proj3d.proj_transform(
x_star[0], x_star[1], y_star,
ax.get_proj()
)
ax.annotate(
"★",
(x2, y2),
color="C3",
fontsize=14,
ha="center",
zorder=20,
)
plt.tight_layout(h_pad=2.5)
plt.show()