Constrained optimization - SEGO#
[1]:
import numpy as np
from matplotlib import pyplot as plt
from smt_optim.benchmarks.registry import get_problem
from smt_optim.core import Driver, ObjectiveConfig, ConstraintConfig, DriverConfig, Problem
from smt_optim.surrogate_models.smt import SmtAutoModel
from smt_optim.acquisition_strategies import MFSEGO
Importing a test function#
[2]:
problem = get_problem("Branin1")
Constrained 2D optimization#
As with unconstrained optimisation, we must first define the problem. The objective is configured using the ObjectiveConfig data class, and each constraint must be defined using the ConstraintConfig data class.
[3]:
obj_config = ObjectiveConfig(
[problem.objective[-1]],
type="minimize",
surrogate=SmtAutoModel,
)
# configure the constraint
cstr_config = ConstraintConfig(
[problem.constraints[0][-1]],
type="less", # set the constraint type (less, greater or equal)
value=0.0, # g(x) <= 0
surrogate=SmtAutoModel, # set which GP to model this constraint
)
prob_definition = Problem(
obj_configs=[obj_config],
design_space=problem.bounds, # problem bounds
cstr_configs=[cstr_config], # list the constraints
)
opt_config = DriverConfig(
max_iter = 10,
nt_init = 6,
verbose = True,
scaling = True,
seed=42,
)
driver = Driver(prob_definition, opt_config, MFSEGO)
state = driver.optimize()
iter budget fmin fidelity gp_time acq_time
1 7 6.86090e+01 1 0.609 0.552
2 8 1.44188e+01 1 0.565 0.289
3 9 7.18465e+00 1 0.526 0.205
4 10 6.94315e+00 1 0.538 0.218
5 11 6.94315e+00 1 0.476 0.357
6 12 6.94314e+00 1 0.452 0.305
7 13 6.94314e+00 1 0.449 0.377
8 14 6.94314e+00 1 0.447 0.245
9 15 6.94314e+00 1 0.433 0.218
10 16 6.94314e+00 1 0.410 0.250
The get_best_sample class method enables us to obtain the optimal sample within a specific constraint tolerance by passing the ctol argument. The code snippet below demonstrates how imposing a stricter tolerance constraint can affect the best sample found.
[4]:
tolerances = [1.2e-1, 1e-4]
for tol in tolerances:
sample = state.get_best_sample(ctol=tol)
print(sample)
======= sample data =======
x = [0.12899267 0.74173099]
obj = [2.00404723]
cstr = [0.10432214]
eval_time = [3.05403955e-06 4.44008037e-07]
------- meta data -------
iter = 0
budget = 2
fidelity = 0
===========================
======= sample data =======
x = [1. 0.20019954]
obj = [6.94314066]
cstr = [-0.00019954]
eval_time = [6.89597800e-06 1.20600453e-06]
------- meta data -------
iter = 10
budget = 16
fidelity = 0
===========================
[5]:
# get the best sample in the dataset
sample = state.get_best_sample(ctol=1e-4)
xt = state.dataset.export_as_dict()["x"]
X = np.linspace(0, 1, 201)
XX, YY = np.meshgrid(X, X)
data = np.vstack((XX.ravel(), YY.ravel())).T
z = np.empty(data.shape[0])
c = np.empty(data.shape[0])
for i in range(data.shape[0]):
z[i] = problem.objective[-1](data[i, :])
c[i] = problem.constraints[0][-1](data[i, :])
Z = z.reshape(XX.shape)
C = c.reshape(XX.shape)
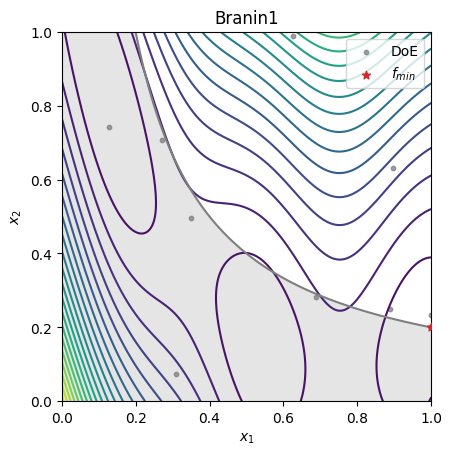
fig, ax = plt.subplots()
ax.set_title(problem.name)
ax.contour(XX, YY, Z, levels=20)
ax.contourf(XX, YY, np.where(C <= 0, np.nan, C), levels=0, colors="C7", alpha=0.20)
ax.contour(XX, YY, C, levels=[0], colors="C7")
ax.scatter(xt[:, 0], xt[:, 1], 10, color="C7", alpha=0.75, label="DoE")
ax.scatter(sample.x[0], sample.x[1], c="C3", marker="*", label=r"$f_{min}$", zorder=10)
ax.set_xlabel(r"$x_1$")
ax.set_ylabel(r"$x_2$")
ax.legend()
ax.set_aspect("equal")
plt.show()