Multi-fidelity optimization#
This page demonstrates how to minimize multi-fidelity problems. The purpose of multi-fidelity optimization is to use cheaper approximations to reduce the budget required for convergence.
This page assumes that the user is already familiar with unconstrained and constrained optimization.
import numpy as np
import matplotlib.pyplot as plt
from smt_optim import minimize
# utility method to help with plotting 2d functions
from smt_optim.utils.plot_2d import get_plot2d_data
Unconstrained multi-fidelity optimization#
Defining the test problem#
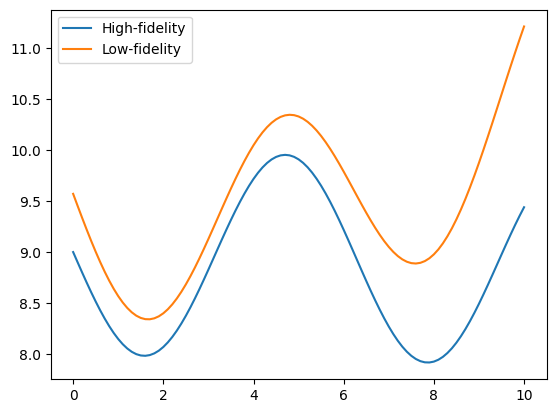
The following example illustrates how to optimize the Sasena 2002 multi-fidelity test function. The problem is defined by a high-fidelity (true) function, and a low-fidelity function which will be used as an approximation. Both functions can be expressed mathematically as:
The objective will be to minimize the high-fidelity function with respect to a bounded domain. It can be expressed as:
The cell below defines the low- and high-fidelity objective functions that should be minimized.
# high-fidelity function
def sasena2002_hf(x):
return -np.sin(x)-np.exp(x/100)+10
# low-fidelity function
def sasena2002_lf(x):
return sasena2002_hf(x)+0.3+0.03*(x-3)**2
bounds = np.array([
[0, 10]
])
x_valid = np.linspace(bounds[0, 0], bounds[0, 1], 101)
y_hf = sasena2002_hf(x_valid)
y_lf = sasena2002_lf(x_valid)
fig, ax = plt.subplots()
ax.plot(x_valid, y_hf, label="High-fidelity")
ax.plot(x_valid, y_lf, label="Low-fidelity")
ax.legend()
plt.show()
Starting the optimization#
This example uses the minimize method to initiate the multi-fidelity Bayesian optimization. When dealing with multi-fidelity optimization, two arguments must be passed:
objective: the function to minimize;design_space: the function boundary;costs: the sampling cost of each fidelity level.
Note
The objective argument requires a list of callables. These callables and their associated sampling costs must be ordered by level of fidelity, from lowest to highest.
Moreover, we can also pass the following optional arguments:
max_iter: the maximum number of iterations before the program stops;driver_kwargs: kwargs to pass to the optimization driver. Thent_initparameter sets the number of high-fidelity samples should be in the initial DoE. In our case,nt_initis set to three samples. Theseedparameter makes this example reproducible.
Note
The initial multi-fidelity DoE is generated using SMT's Nested Latin Hypercube Sampling, meaning it will have the following properties:
- Samples will be nested;
- Each lower-fidelity level will have twice as many samples as the next higher-fidelity level.
The minimize method returns a State object that contains the design of experiment (DoE) with all the low- and high-function samples. Using these samples, we can identify the best sampled function value found during the optimization process.
state = minimize(
[sasena2002_lf, sasena2002_hf], # in increasing order of fidelity
bounds,
costs=[0.2, 1.], # in increasing order of fidelity
max_iter=10,
driver_kwargs={
"nt_init": 3, # number of sample in the initial Design of Experiment (DoE)
"seed": 0, # makes this example reproducible
}
)
iter budget fmin rscv fidelity gp_time acq_time
1 4.400 8.07060e+00 0.000e+00 1 0.134 0.160
2 4.600 8.07060e+00 0.000e+00 1 0.118 0.151
3 5.800 7.98975e+00 0.000e+00 2 0.111 0.165
4 6.000 7.98975e+00 0.000e+00 1 0.223 0.312
5 6.200 7.98975e+00 0.000e+00 1 0.211 0.263
6 7.400 7.91824e+00 0.000e+00 2 0.198 0.288
7 8.600 7.91824e+00 0.000e+00 2 0.167 0.266
8 9.800 7.91824e+00 0.000e+00 2 0.177 0.297
9 11.000 7.91824e+00 0.000e+00 2 0.166 0.249
10 11.200 7.91824e+00 0.000e+00 1 0.180 0.280
The get_best_sample class method accepts a fidelity argument that specifies the level from which to retrieve the best sample. By default, it fetches the best sample from the highest fidelity level.
print("####### Best low-fidelity sample #######")
best_sample = state.get_best_sample(fidelity=0)
print(best_sample)
print("####### Best high-fidelity sample #######")
best_sample = state.get_best_sample()
print(best_sample)
####### Best low-fidelity sample #######
======= sample data =======
x = [1.68378209]
obj = [8.34136862]
cstr = []
eval_time = [1.25820006e-05]
------- meta data -------
iter = 2
budget = 4.6000000000000005
fidelity = 0
rscv = 0.0
===========================
####### Best high-fidelity sample #######
======= sample data =======
x = [7.86479479]
obj = [7.91823506]
cstr = []
eval_time = [3.51500057e-06]
------- meta data -------
iter = 8
budget = 9.8
fidelity = 1
rscv = 0.0
===========================
Plotting the results#
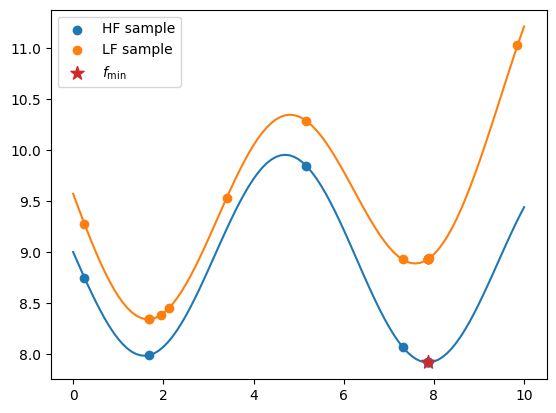
The code snippet below exports all the evaluated low- and high-fidelity design points, as well as their corresponding objective values. It plots these values against their respective fidelity functions. The best sample is marked with a star.
When exporting the dataset using the export_as_dict class method, use the fidelity array to distinguish between different fidelity levels, as demonstrated below.
data = state.dataset.export_as_dict()
# retrieves the fidelity ID of each sample
fidelity = data["fidelity"]
# retrieves the low fidelity DoE
x_doe_lf = data["x"][fidelity == 0]
y_doe_lf = data["obj"][fidelity == 0]
# retrieve the high fidelity DoE
x_doe_hf = data["x"][fidelity == 1]
y_doe_hf = data["obj"][fidelity == 1]
fig, ax = plt.subplots()
# plots the HF function and samples
ax.plot(x_valid, y_hf)
ax.scatter(x_doe_hf, y_doe_hf, label="HF sample", color="C0")
# plots the LF function and samples
ax.plot(x_valid, y_lf)
ax.scatter(x_doe_lf, y_doe_lf, label="LF sample", color="C1")
# plots the best (HF) sample
ax.scatter(best_sample.x, best_sample.obj, label=r"$f_\min$", marker="*", color="C3", s=100, zorder=50)
ax.legend()
plt.show()
Constrained multi-fidelity optimization#
Defining the test problem#
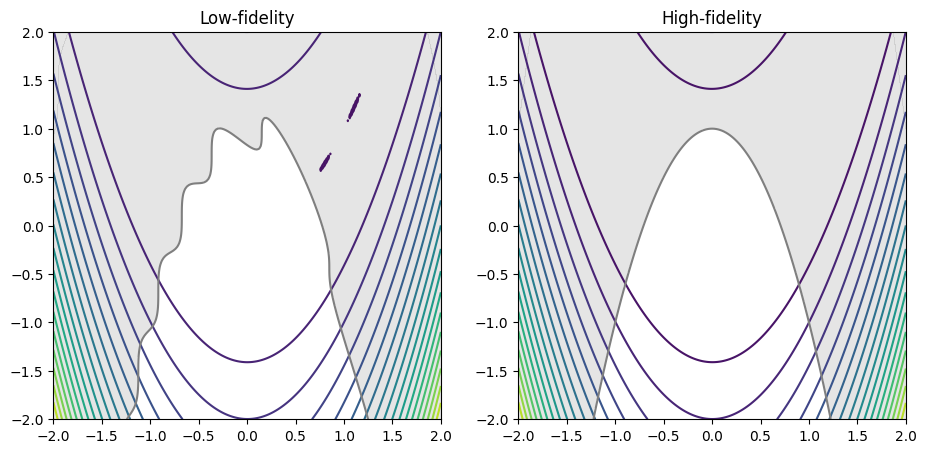
The following example demonstrates how to minimize the multi-fidelity constrained Rosenbrock test problem. The high- and low fidelity objective and constraint functions are as follows:
The objective will be to minimize the high fidelity objective function with respect to the high-fidelity inequality constraint:
The cell below defines the low- and high-fidelity objective and constraint function.
Note
The minimize method requires all quantities of interest (QoI) (e.g.: objectives and constraints) to have the same number of fidelities.
# defines the high-fidelity objective function
def rosenbrock_hf(x):
return (1 - x[0])**2 + 100*(x[1] - x[0]**2)**2
# defines the low-fidelity objective function
def rosenbrock_lf(x):
return rosenbrock_hf(x) + 0.1*np.sin(10*x[0] + 5*x[1])
# defines the high-fidelity constraint function
def constraint_hf(x):
return -(-x[0]**2 - (x[1] - 1)**1/2)
# defines the low-fidelity constraint function
def constraint_lf(x):
return constraint_hf(x) - 0.1*np.sin(10*x[0] + 5*x[1])
# defines the problem boundary
bounds = np.array([
[-2, 2],
[-2, 2]
])
# plots the low- and high-fidelity test problem
XX, YY, Z_hf = get_plot2d_data(rosenbrock_hf, bounds, 201)
_, _, Z_lf = get_plot2d_data(rosenbrock_lf, bounds, 201)
_, _, G_hf = get_plot2d_data(constraint_hf, bounds, 201)
_, _, G_lf = get_plot2d_data(constraint_lf, bounds, 201)
fig, ax = plt.subplots(1, 2, figsize=(11, 7))
ax[0].set_title("Low-fidelity")
ax[0].contour(XX, YY, Z_lf, levels=20)
ax[0].contourf(XX, YY, np.where(G_lf <= 0, np.nan, G_lf), levels=0, colors="C7", alpha=0.20)
ax[0].contour(XX, YY, G_lf, levels=[0], colors="C7")
ax[0].set_aspect("equal")
ax[1].set_title("High-fidelity")
ax[1].contour(XX, YY, Z_hf, levels=20)
ax[1].contourf(XX, YY, np.where(G_hf <= 0, np.nan, G_hf), levels=0, colors="C7", alpha=0.20)
ax[1].contour(XX, YY, G_hf, levels=[0], colors="C7")
ax[1].set_aspect("equal")
plt.show()
Starting the optimization#
The minimize method is called to begin the constrained multi-fidelity Bayesian optimization. Three arguments must be passed when dealing with constrained multi-fidelity optimization:
objective: the function to minimize;design_space: the function boundary;costs: the sampling cost of each fidelity level;constraints: the constraint definitions.
Note
The objective and the constraints callables, as well as the sampling costs must be ordered in increasing level of fidelity and must have the identical number of callables.
Moreover, we can also pass the following optional arguments:
max_iter: the maximum number of iterations before the program stops;max_budget: the maximum budget before the program stops. The budget is computed with the sampling costs;driver_kwargs: kwargs to pass to the optimization driver. Theseedparameter makes this example reproducible;strategy_kwargs: kwargs to pass to the acquisition strategy.
Note
The strategy_kwargs depend on the AcquisitionStrategy selected by the minimize method. For a multi-fidelity problem (constrained or unconstrained), the MFSEGO acquisition strategy is selected. The valid arguments can be consulted via the API reference.
The minimize method returns a State object that contains the design of experiment (DoE) with all the low- and high-function samples. Using these samples, we can identify the best sampled function value found during the optimization process.
# defines the inequality constraint
constraint = [
{
"fun": [constraint_lf, constraint_hf], # in increasing order of fidelity
"upper": 0., # equivalent to: g(x) <= 0
}
]
state = minimize(
[rosenbrock_lf, rosenbrock_hf], # in increasing order of fidelity
bounds,
costs=[0.2, 1.],
constraints=constraint,
max_iter=30,
max_budget=15.,
# arguments passed to the optimization driver
driver_kwargs={
"seed": 0
},
# arguments passed to the acquisition strategy
strategy_kwargs={
"fidelity_crit": "pessimistic",
"sp_method": "Cobyla",
"sp_tol": 1e-4
}
)
iter budget fmin rscv fidelity gp_time acq_time
1 7.200 2.64887e+00 0.000e+00 1 0.804 1.917
2 7.400 2.64887e+00 0.000e+00 1 0.761 1.434
3 7.600 2.64887e+00 0.000e+00 1 0.699 1.485
4 7.800 2.64887e+00 0.000e+00 1 0.760 1.383
5 8.000 2.64887e+00 0.000e+00 1 0.702 2.299
6 8.200 2.64887e+00 0.000e+00 1 0.756 2.276
7 8.400 2.64887e+00 0.000e+00 1 0.696 2.596
8 8.600 2.64887e+00 0.000e+00 1 0.719 2.708
9 8.800 2.64887e+00 0.000e+00 1 0.691 2.475
10 9.000 2.64887e+00 0.000e+00 1 0.697 2.457
iter budget fmin rscv fidelity gp_time acq_time
11 9.200 2.64887e+00 0.000e+00 1 0.725 2.303
12 9.400 2.64887e+00 0.000e+00 1 0.586 1.840
13 9.600 2.64887e+00 0.000e+00 1 0.714 1.892
14 10.800 2.64887e+00 0.000e+00 2 0.636 1.784
15 12.000 2.78736e-01 0.000e+00 2 0.661 1.315
16 12.200 2.78736e-01 0.000e+00 1 0.607 2.422
17 13.400 1.79294e-01 0.000e+00 2 0.628 2.291
18 14.600 1.78500e-01 2.982e-05 2 0.592 2.645
19 15.800 1.78479e-01 1.487e-05 2 0.597 2.507
Plotting the results#
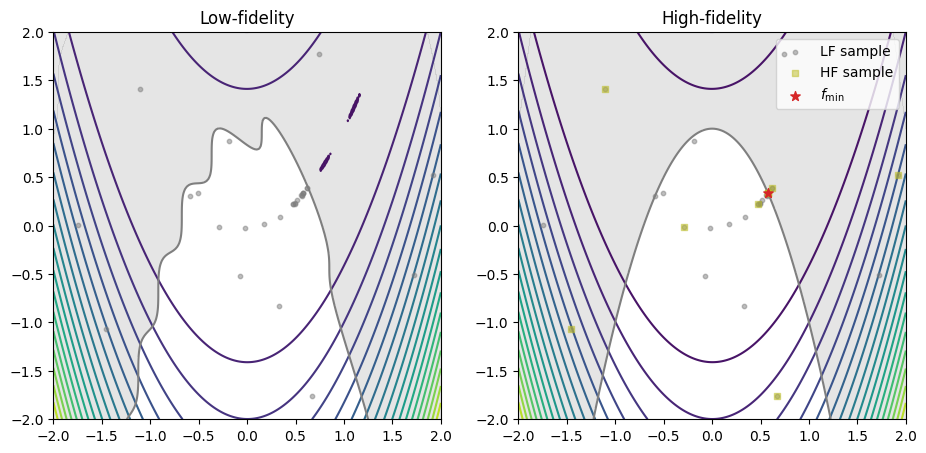
The code snippet below exports all the evaluated low- and high-fidelity design points, as well as their corresponding objective values, plotting them against their respective fidelity function. The best sample is marked with a star.
When exporting the dataset using the export_as_dict class method, use the fidelity array to distinguish between different fidelity levels, as demonstrated below.
# gets the best sample (minimum feasible objective value)
best_sample = state.get_best_sample()
data = state.dataset.export_as_dict()
# retrieves the fidelity ID of each sample
fidelity = data["fidelity"].ravel()
# retrieves the low fidelity DoE
x_doe_lf = data["x"][fidelity == 0, :]
# y_doe_lf = data["obj"][fidelity == 0]
# retrieve the high fidelity DoE
x_doe_hf = data["x"][fidelity == 1, :]
# y_doe_hf = data["obj"][fidelity == 1]
fig, ax = plt.subplots(1, 2, figsize=(11, 7))
ax[0].set_title("Low-fidelity")
ax[0].contour(XX, YY, Z_lf, levels=20)
ax[0].contourf(XX, YY, np.where(G_lf <= 0, np.nan, G_lf), levels=0, colors="C7", alpha=0.2)
ax[0].contour(XX, YY, G_lf, levels=[0], colors="C7")
ax[0].set_aspect("equal")
ax[0].scatter(x_doe_lf[:, 0], x_doe_lf[:, 1], s=10, color="C7", alpha=0.5, label="LF sample", zorder=50)
ax[1].set_title("High-fidelity")
ax[1].contour(XX, YY, Z_hf, levels=20)
ax[1].contourf(XX, YY, np.where(G_hf <= 0, np.nan, G_hf), levels=0, colors="C7", alpha=0.2)
ax[1].contour(XX, YY, G_hf, levels=[0], colors="C7")
ax[1].set_aspect("equal")
# scatters the LF samples
ax[1].scatter(x_doe_lf[:, 0], x_doe_lf[:, 1], s=10, color="C7", alpha=0.5, label="LF sample", zorder=50)
# scatters the HF samples
ax[1].scatter(x_doe_hf[:, 0], x_doe_hf[:, 1], s=20, color="C8", alpha=0.5, marker="s", label="HF sample", zorder=60)
# plots the best sample
ax[1].scatter(best_sample.x[0], best_sample.x[1], s=50, color="C3", marker="*", label=r"$f_\min$", zorder=100)
ax[1].legend(loc="upper right")
plt.show()
Many multi-fidelity levels optimization#
The minimize method supports more than two fidelity levels. An example coming soon.
